Let’s kick off this discussion with a simple calculation about the length of DNA in a single diploid human cell. We know according to Watson and Crick model of double helix DNA, each two base pairs has a distance of 3.4 A. There are approximately 6.4 billion base pairs (bp) in a human cell. So, the total length of a DNA (if all the DNA of 23 pairs are assumed to be joined end to end) molecule is about 2.2 meter.
So a typical human cell contains about 2.2 m of DNA. Now the problem arises when we think about placing this large DNA strand in a very microscopic nucleus of about 10 micrometer in diameter. To solve this problem, it is obvious that the DNA molecule needs to be folded many times inside the nucleus. And this thing actually does happen.
To support the DNA’s wrapping and packaging inside the nucleus, many protein molecules assist which are mainly histone proteins. Experimentally it was seen that the ratio of DNA and histone proteins in the chromosome is 1:1.
Many theories have been proposed regarding the packaging nature of the DNA and histone proteins together. One of them is Nucleosome model which is studied below.
Discoverer
P. Odet et. al (1975).
What is a nucleosome?
Nucleosome is the fundamental repeating unit of a chromosome organization in eukaryotic cells composed of a double stranded DNA wrapped around a set of 8 histone proteins. So, A nucleosome = A DNA (146 bp) + 8 histone proteins.
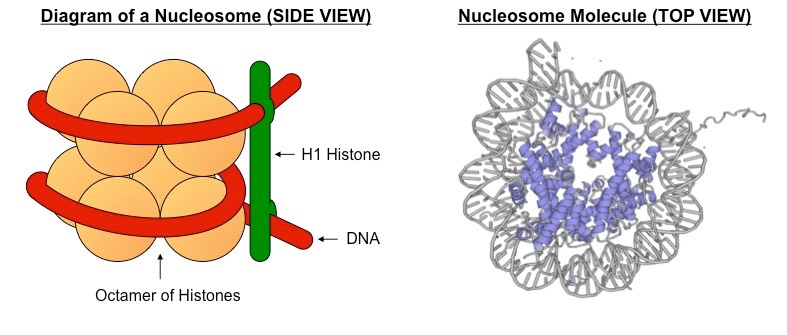
- The wrapped DNA molecule of a nucleosome roughly contains about 146 bp (In the above figure, the red strand of DNA in the left contains 146 bp).
- The DNA around the histone is called core or wrapping DNA (Red colored) which is folded two and a half times.
- The histones are typically of 4 types consisting two of each type. Such as: H2A, H2B, H3, H4.
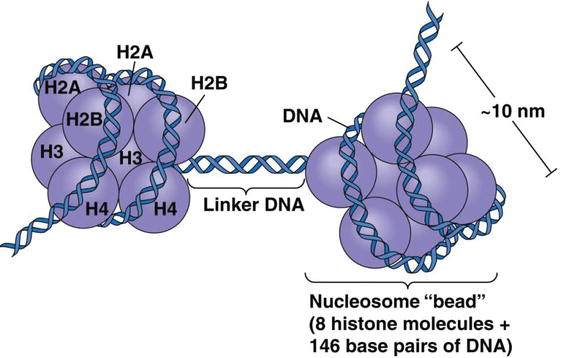
Source: Wikipedia - Microscopically it was observed that each nucleosome has a length of about 100 A (=10 nm) and width 60 A. It is the least packed form of a chromatin fiber.
- The 8 histones are together called Histone octamers.
- As mentioned before, many nucleosomes are repeated in a chromosome molecule. These repeated bead like structures are connected via a linker DNA typically consisting of 54 bp and a H1 histone protein with that DNA molecule.
- So, H1 histone protein binds to the site where DNA enters and exits the nucleosome.
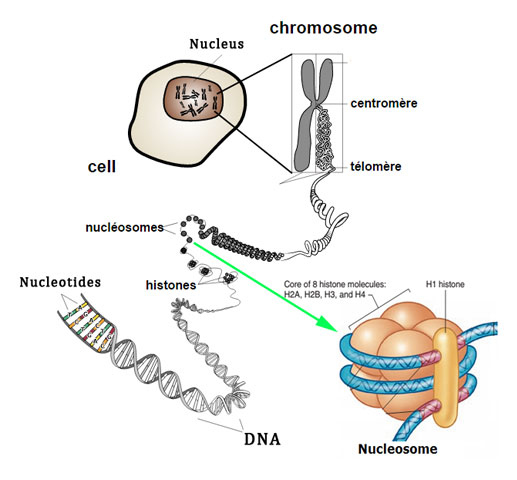
Good to know:
- Nucleosome model is the most accepted model of chromatin organization.
- A chromatin is formed by a DNA molecule and its associate proteins.
- The histone proteins are positively charged basic proteins as they contain basic amino acids such as lysine and arginine which typically contains amino group that may receive protons to become positively charged.
- There are five types of histones in eukaryotes based on structural differences, molecular weight and lysine, arginine ratio.
- Five types of histones: H1, H2A, H2B, H3 and H4.
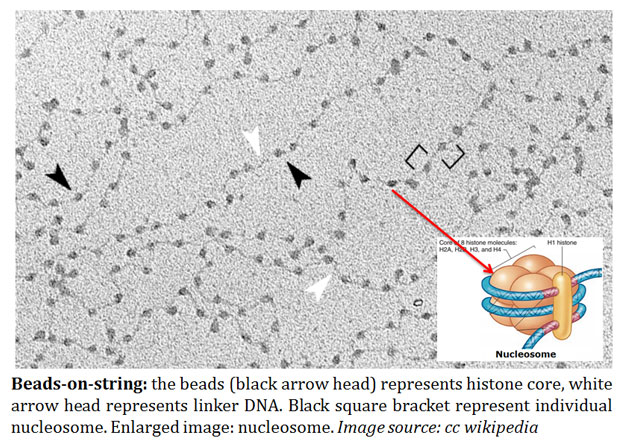
Source:
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
Best Money Earning Website 100$ Day
#1 Top ranking article submission website
- https://www.easybiologyclass.com/nucleosome-model-of-chromosomes-in-eukaryotes-short-notes/
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka





Can you be more specific about the content of your article? After reading it, I still have some doubts. Hope you can help me.