Bacteria (singular: bacterium) are classified as prokaryotes. They are single-celled life/body with a simple core-less internal structure and contain DNA that swims freely in a twisted, thread-like matter called a nucleoid, or in circular parts called plasmids. Bacteria are single-celled microscopic life/organisms that grow in different environments, and these organisms can live in the ocean, soil and inside the human intestine.
Gross Morphology
Some characteristic shapes like (spiral, rods, cocci etc.) and some characteristic multitude/combination like (tetrads, chains, pairs, clusters etc.) arise/arrive in bacteria. They are important to human life and also life on planet Earth but bacteria have nothing to do with human life as living existence can be. Though they are famous for their nature for causing human illness. From tooth decay to the Black Death, there are some favourable/helpful species that are essential for a soundness and recovery health.
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
Best Money Earning Website 100$ Day
#1 Top ranking article submission website
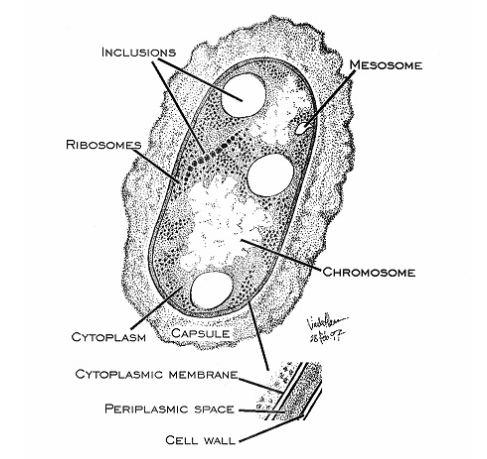
Cell Structure
Prokaryotes have a nucleoid (nuclear body) rather than an enveloped nucleus and do not have cytoplasmic organelles bound to the membrane. The plasma membrane of prokaryotes plays many functions which performed by membrane organelles in eukaryotes. Binary fission results multiplication.
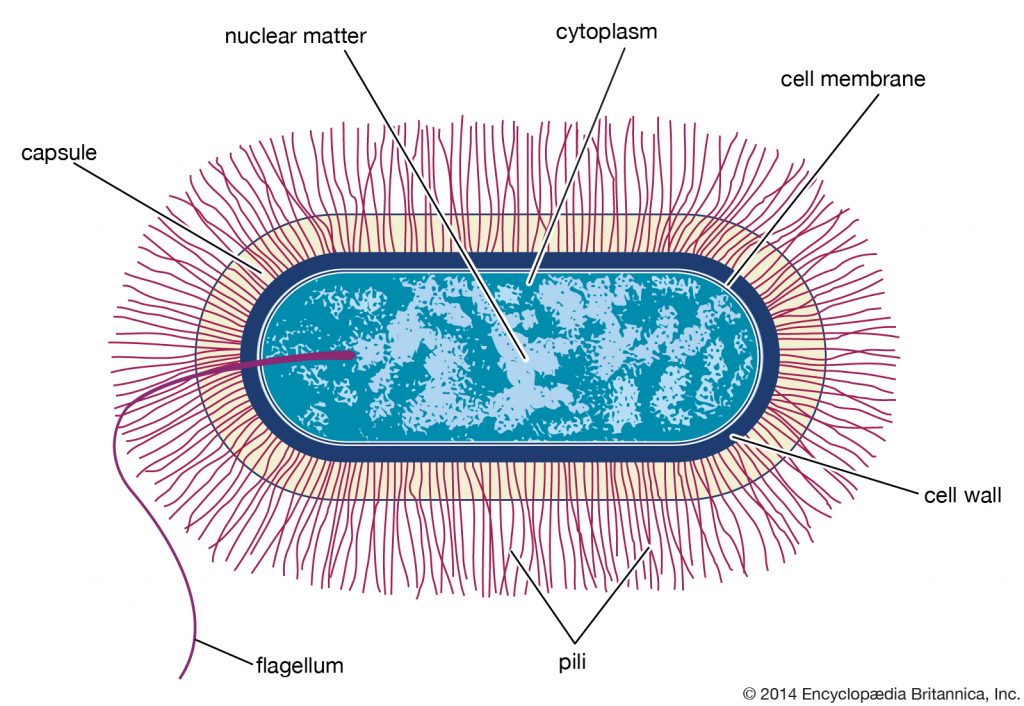
Structurally, there are three architectural regions: The appendages (attachments to the cell surface) in the form of flagella and pili (or fimbriae). A cell envelope consisting of a capsule, a cell wall and a plasma membrane which contains the ribosomes, cell chromosome (DNA) and various kinds of inclusions.
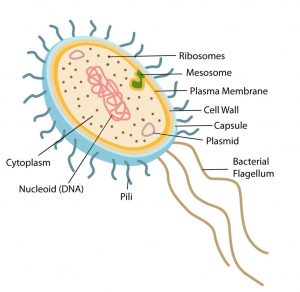
Surface Structures
1. Flagellum (plural: flagella) are helical structure which is composed/fixed of subunits of a protein called flagellin. At the base of the flagellum has a wider region which is called a hook. It is also different in structure than that of the filament. This hook connects the filament to the motor portion of the flagellum called a basal body. And the basal body is anchored in the cell wall and cytoplasmic membrane. There are presence of rings that are surrounded by a pair of proteins called Mot. These proteins actually accelerate the flagellar motor causing rotation of the filament. Another set of proteins called Fli. This Fli proteins works as the motor switch, reversing/overturn the rotation of the flagella in response to intracellular signals.
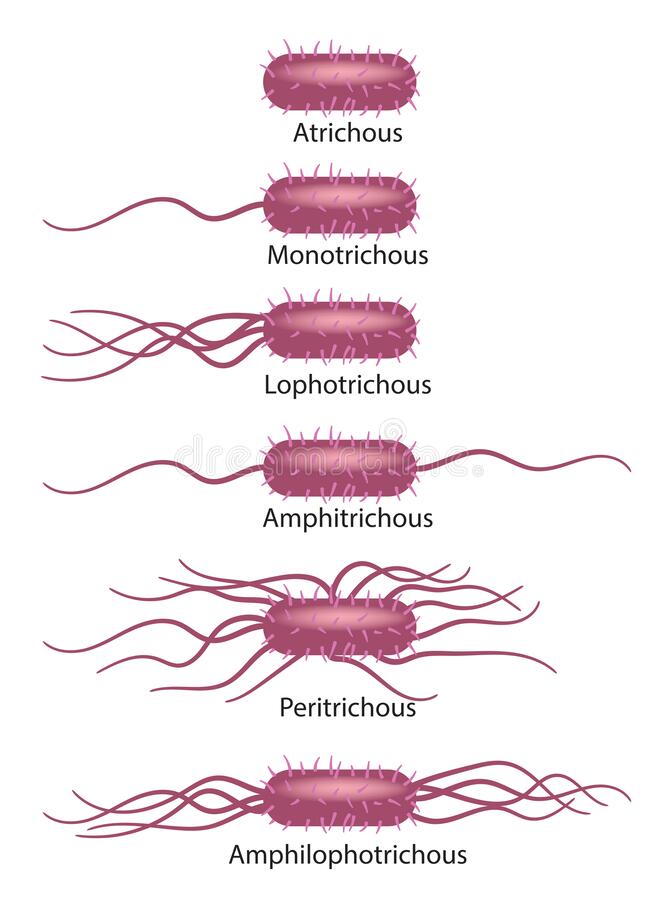
The location and number of flagella are distinguishable for each genus. Four types of flagellar arrangements we usually saw:
- Monotrichous (Mono means one): Single polar flagellum e.g. Vibrio cholerae, Campylobacter spp. (To give a “seagull” appearance polar flagella often in pairs).
- Amphitrichous: Single flagellum at both ends e.g. Alcaligenes faecalis (Note: amphibians live both in water and on land).
- Lophotrichous: Tuft of flagella at one or both ends e.g. Spirilla spp.
- Peritrichous (flagella in periphery): Flagella surrounding the bacterial cell. All the members of family Enterobacteriaceae, if motile have peritrichous flagella. e.g. Salmonella typhi, Escherichia coli, Proteus spp (highly motile organism; shows swarming motility).
2. Pili are hair-like fibers which made up of thousands of protein subunits (pilin, 16–20 k Da). Neisseria spp. Pathogens have long pili (up to 4300 nm in length), is called type IV pili, which protrude from the bacterial surface. Neisseria spp. which is non-pathogenic can express both short and long pili (175–210 nm in length). Type IV pili confer/offer bacterial cell-to-cell interactions and contraction motility – a form of locomotion fueled by the extension and retraction of the pilus filament. This Pili are essential for adhesion to epithelial and endothelial cells and confer tissue tropism. Expression of type IV pili is also required for DNA transformation. During bacterial conjugation the conjugative pili allow the transfer of DNA between bacteria. By analogy with sexual reproduction, they are sometimes called “sexual pili”, because they always allow the exchange of genes by the formation of “mating couples”. Possibly the most studied is the F pilus from Escherichia coli which is encoded by the fertility factor or the F plasmid.
Important Chemical Components of Surface Structures
Cell Wall Peptidoglycans: Both gram-positive and gram-negative bacteria have their cell wall peptidoglycans. Which give the cell its own characteristic shape. It also offers the cell mechanical protection. For prokaryotic organisms peptidoglycans are unique and consist of a glycan backbone of glucosamine and muramic acid (both N-acetylated) and peptide chains that are partially cross-linked or strongly cross-linked with bridges in gram-positive bacteria (e.g. Staphylococcus aureus) B. (Escherichia coli). The targets for b-lactam antibiotics are the cross-linking transpeptidase enzymes .
Teichoic Acids: Teichoic acids are mainly polyol phosphate polymers which have a strong negative charge. Teichoic acids are covalently attached to peptidoglycan in several Gram positive bacteria. They are strongly antigenic, but usually absent in Gram negative bacteria.
Lipoteichoic Acids: Lipoteichoic acids as membrane teichoic acids are anchored in the cytoplasmic membrane and they are polymers of amphiphitic glycophosphates with lipophilic glycolipid. They are adhesins, cytotoxic and antigenic (eg, Streptococcus pyogenes).
Lipopolysaccharides: Lipopolysaccharide (endotoxin) is one of the major components of outer membrane of gram-negative bacteria, a complex molecule which consisting of a lipid A anchor, carbohydrate chains and a polysaccharide core. Polysaccharide chains of the sugar confer serological specificity.
Wall-Less Forms: Two groups of bacteria without cell wall peptidoglycans are the Mycoplasma species, which have a membrane structure, surface, and the L-forms, which arise from either Gram-negetive or Gram-positive bacterial cells that have lost their capability to produce the peptidoglycan structures.
The Nucleoid
An important advance in recent years has been the discovery that the circular DNA of the bacterial chromosome is known as the “nucleoid”. The nucleoid (i.e. nuclear-like) is an irregularly shaped region within a prokaryote’s cell that contains all or most of the genetic material. In contrast to the nucleus of a eukaryotic cell, it is not surrounded by a nuclear membrane. The genome of prokaryotic organisms is generally a circular, double-stranded piece of DNA, of which multiple copies can exist at any one time. The length of a genome varies widely, but is generally at least a few million base pairs.
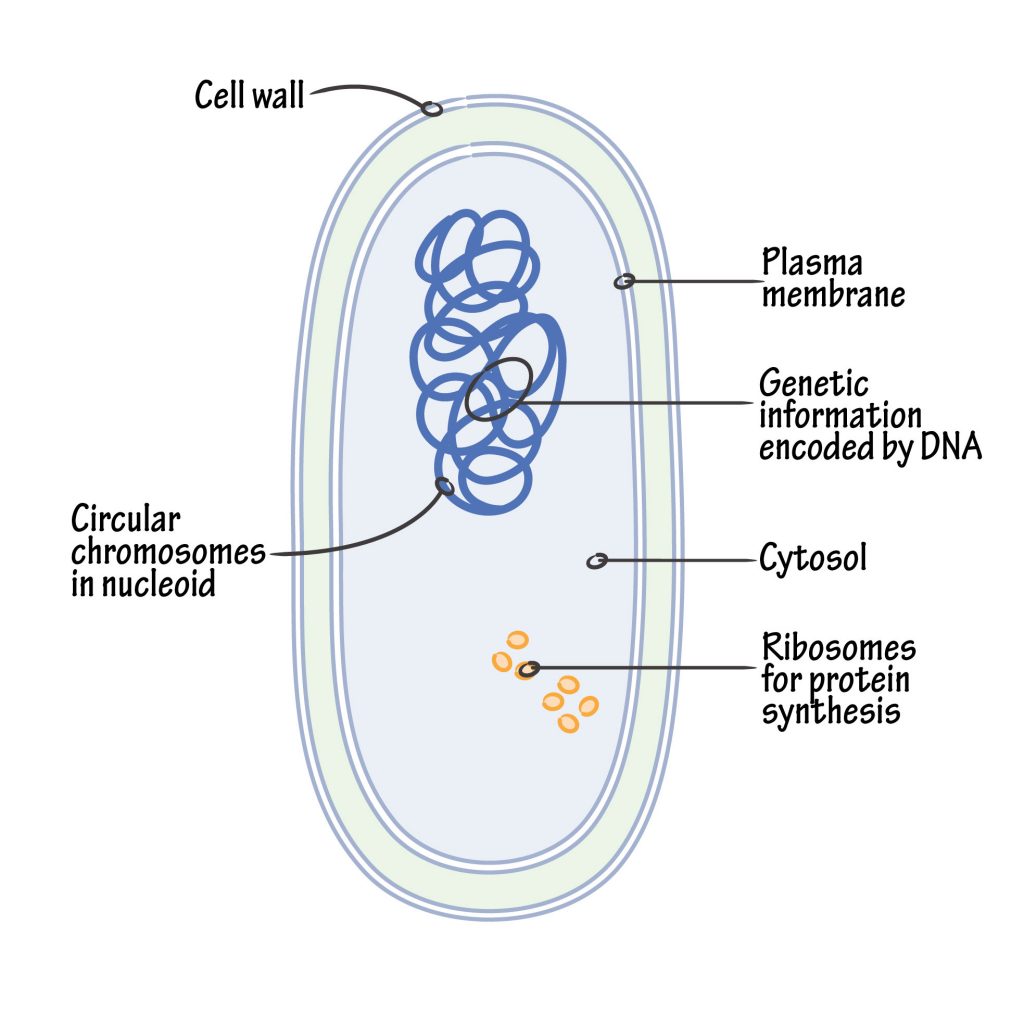
Bacterial chromatin does not contain basic histone proteins, but low molecular weight polyamines and magnesium ions can serve a similar function as eukaryotic histones. Despite the differences between prokaryotic and eukaryotic DNA, prokaryotic DNA from cells infected with bacteriophage, when visualized by electron microscopy, has a pearly, condensed appearance not dissimilar to that of eukaryotic chromatin.
Cytoplasmic Structures
Plasma Membrane: The bacterial plasma-membrane is determined/composed of primarily protein and phospholipid (about 3:1). It plays many functions, including biosynthesis, transport and energy transduction.
Organelles: The bacterial cytoplasm is tightly packed with 70S ribosomes. Other granules represent metabolic reserves (poly-β-hydroxybutyrate, polysaccharide, polymetaphosphate and metachromatic granules).
Endospores: Clostridium and Bacillus species can produce endospores: dehydrated dormant, heat-resistant cells that are formed intra-cellularly and contain single genome and all important metabolic machines. A complex protective layer surrounds mostly the endospore.
Capsules
Capsules are the outermost structures of fungal and bacterial cells. Capsules that form the outermost layer of the bacterial cell and surround it with a relatively thick layer of viscous gel. Capsules can be up to 10 µm thick. Some organisms do not have a well-defined capsule, but they have loose, amorphous layers of mucus outside the cell envelope or wall. The capsules protects microbial cell from immune recognition and killing during infection of mammalian hosts. Ensured/certain bacteria (e.g. Streptococcus pyogenes and Anthracis) produce only one capsule structure, while many other bacteria express their capsules with great immunological, biochemical and structural diversity within the same species. This diversity is always accelerate by immune selection from the mammalian hosts. The genes for capsule synthesis are typically clustered in a single location on the bacterial chromosome. The number of genes associated with capsule synthesis is in the range. Pointed out capsule polymers also have important/essential industrial uses.
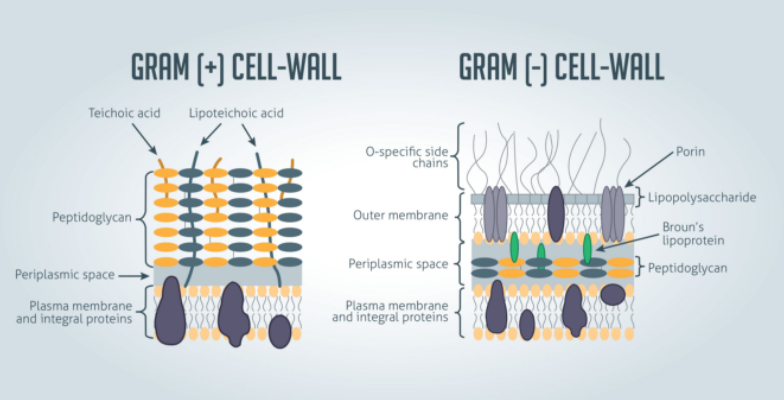
Cell Wall and Gram-Negative and Gram-Positive Cell Envelope
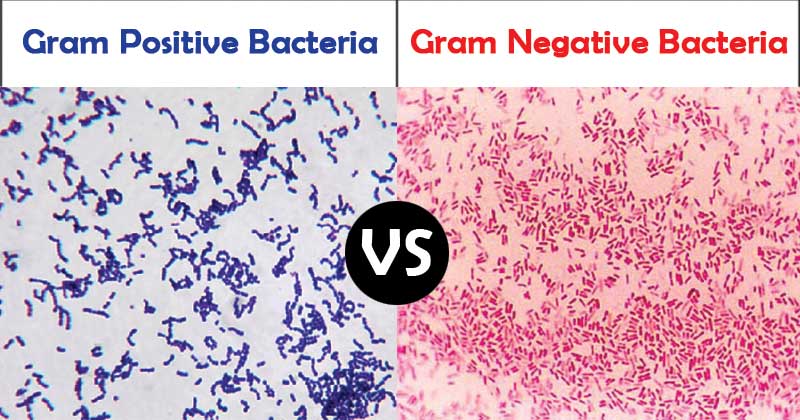
Gram staining roughly distinguishes bacteria into gram-positive and gram-negative groups; Some organisms are entirely gram-variable. Gram-positive and gram-negative organisms differ drastically in the organization of the structures outside the plasma membrane, but below the capsule.
Gram-negative bacteria are surrounded by a thin peptidoglycan cell wall, which is itself surrounded by an outer membrane that contains lipopolysaccharide. Gram-positive bacteria do not have an outer membrane, but are surrounded by peptidoglycan layers that are many times thicker than the gram-negative bacteria. The following figure shows the differences in the structure of gram-positive and gram-negative bacteria.
The two main features that lead to the different visualization properties of Gram-positive and Gram-negative species are the thickness of the peptidoglycan layer and the presence or absence of the outer lipid membrane. This is because the wall structure affects the cell’s ability to retain the crystal violet stain used in the Gram staining process, which can then be visualized under a light microscope.
Accessory Wall Polymers
In addition to the main cell wall polymers, the walls of certain gram-positive bacteria have polysaccharide molecules that are bound to the peptidoglycan. For example, the C-polysaccharide of Streptococci offers/confers group specificity. Acid polysaccharides which bound to the peptidoglycan are called teichuronic acids. Mycobacteria have peptidoglycolipids, glycolipids, and waxes that are associated with the cell wall.
Intracellular Components
1. Plasma Membranes
Bacterial plasma membranes, the functional equivalents of eukaryotic plasma membranes, are variously referred to as cytoplasmic, protoplast, or (in Gram-negative organisms) inner membranes. Similar in their overall dimensions and appearance in thin sections like biomembranes from eukaryotic cells, they consist mainly of proteins and lipids (mainly phospholipids). The protein-to-lipid ratio of bacterial plasma membranes is approximately 3: 1 and is close to that of mitochondrial membranes. In contrast to eukaryotic cell membranes, the bacterial membrane (with the exception of Mycoplasma species and certain methylotrophic bacteria) does not contain sterols, and bacteria lack the enzymes necessary for sterol biosynthesis.
2. Mesosomes
Thin sections of gram-positive bacteria reveal the presence of vesicular or tubular-vesicular membrane structures called mesosomes, which appear to be formed by intussusception of the plasma membrane. These structures are much more pronounced in gram-positive organisms than in gram-negative organisms. At one time, the mesosomal vesicles were thought to correspond to bacterial mitochondria; However, many other membrane functions have also been ascribed to the mesosomes. There is currently no satisfactory evidence that they have a unique biochemical or physiological function. In fact, electron microscopic studies have shown that the mesosomes, such as those normally seen in thin sections, can be caused by membrane disorders and fixation artifacts. However, there is no general agreement about this theory, and some evidence suggests that mesosomes may be related to events in the cell division cycle.
Other Intracellular Components
In addition to the nucleoid and cytoplasm (cytosol), the intracellular compartment of the bacterial cell is densely packed with ribosomes of the 70S type. These ribonucleoprotein particles, which have a diameter of 18 nm, are not arranged on a membranous rough endoplasmic reticulum as they are in eukaryotic cells. Other granular inclusions randomly distributed in the cytoplasm of various species include metabolic reserve particles such as poly-β-hydroxybutyrate (PHB), polysaccharide and glycogen-like granules, and polymetaphosphate or metachromatic granules.
Reproduction
Most bacteria multiply by a process which called binary fission. By this process, one single bacterial cell, called the “mother,” which makes a copy of its DNA and then grows by duplicating it’s cellular content. That cell then divides, expelling the duplicated material and creating two identical “daughter” cells.
Some bacterial species, such as cyanobacteria and firmicutes, reproduce through budding. In this case, the daughter cell grows as a branch of the parent. It starts out as a small bump, grows to the same size as its parent, and then splits.
The DNA found in parents and offspring after binary fission or budding is exactly the same. Therefore, bacterial cells introduce the variations in their genetic material by integrating additional DNA, often from their environment, into their genome. This is also known as horizontal gene transfer. The resulting genetic variation ensures that bacteria can adapt and survive as their environmental changes. There are three ways by which horizontal gene transfer occurs: transformation, transduction, and conjugation.
Transformation is the most common horizontal gene transfer process and occurs when short pieces of DNA are exchanged between donors and recipients. Transduction, which generally only occurs between closely related bacteria, requires donor and recipient to transfer DNA by sharing cell surface receptors.
Bacteria in human health and disease
Reference
This Article is based on by the lecture of Dr. Mohammad Abdul karim, Professor, University of Dhaka .
Some info and pictures have been added by Author and from Wikipedia.
Reference used for info: Cambridge International Biology Course Book and other resources.
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka




Precise and informative article
Your article gave me a lot of inspiration, I hope you can explain your point of view in more detail, because I have some doubts, thank you.
Thank you for your sharing. I am worried that I lack creative ideas. It is your article that makes me full of hope. Thank you. But, I have a question, can you help me?
Can you be more specific about the content of your article? After reading it, I still have some doubts. Hope you can help me. https://accounts.binance.com/en-ZA/register?ref=JHQQKNKN
Can you be more specific about the content of your article? After reading it, I still have some doubts. Hope you can help me.