The process of transcription is very similar in prokaryotes and eukaryotes, but there are major differences in the relation between the transcript and the mRNA used for polypeptide synthesis in eukaryotes.
In prokaryotes, the immediate product of transcription (the primary transcript) is mRNA; by contrast, the primary transcript (also called pre-mRNA) in eukaryotes must be converted into mRNA.
This conversion, which is called RNA processing, usually consists of two types of events:
- Modification of the ends (Addition of 5′ cap and poly A tail in 3′ end).
- Excision of untranslated sequences embedded within coding sequences.
Good to know
- In bacterial cells, transcription and translation take place simultaneously.
Modification of the ends
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
Best Money Earning Website 100$ Day
#1 Top ranking article submission website
Addition of 5′ cap
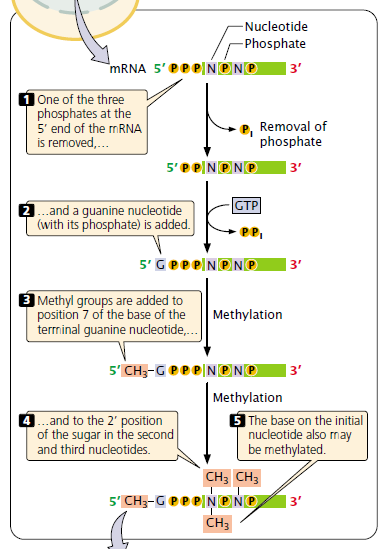 Each end of a eukaryotic transcript is processed. The 5′ end is altered by the addition of a modified guanosine, 7- methyl guanosine, in an uncommon 5′-to-5′ (instead of 3′-to-5′) linkage; this terminal group is called a cap.
Each end of a eukaryotic transcript is processed. The 5′ end is altered by the addition of a modified guanosine, 7- methyl guanosine, in an uncommon 5′-to-5′ (instead of 3′-to-5′) linkage; this terminal group is called a cap.
The process of addition: This capping consists of the addition of an extra nucleotide at the 5′ end of the mRNA and methylation by the addition of a methyl group (CH3) to the base in the newly added neucleotide and (may be) to the 2’–OH group of the sugar of one or more nucleotides at the 5′ end.
So, for easy understanding:
- Addition of an extra nucleotide at the 5′ end of the mRNA.
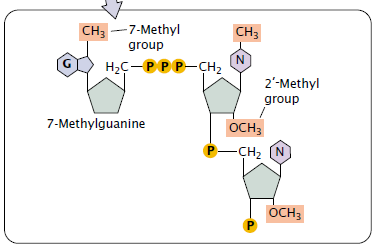
- Methylation to the base (at position 7. so, 7-methyl) in the newly added neucleotide.
- Methylation (may be) to the 2’–OH group of the sugar of one or more nucleotides at the 5′ end (second and third Sugars in the figure below).
Reason: The 5′ cap is necessary for the mRNA to bind with the ribosome to begin protein synthesis. Its presence also increases the stability of mRNA and influences the removal of introns.
Note that
- Cap binding proteins recognize the cap and attach to it; a ribosome then binds to these proteins and moves downstream along the mRNA until the start codon is reached and translation begins.
- Why are 3 phosphates in the 5′ end?
The mRNA is transcribed in 5′ to 3′ direction. And the nucleotides are added from 5′ to downstream in a fashion of monophosphate but they come as tri-phosphate at first (phosphate breakage gives the process a huge deal of energy). Now as the first three phosphates are not cleaved from the first ribonucleoside triphosphate (unlike other triphosphates) in the transcription reaction, the 5′ end has 3 phosphates added.
The Addition of the Poly A tail
The 3′ terminus of a eukaryotic mRNA molecule is usually modified by the addition of a polyadenosine sequence (the poly-A tail) of as many as 50 to 250 nucleotides. These nucleotides are not encoded in the DNA but are added after transcription in a process termed poly-adenylation.
Process of addition: Processing of the 3′ end of pre-mRNA requires sequences both upstream and downstream of the cleavage site. The consensus sequence AAUAAA is usually from 11 to 30 nucleotides upstream of the cleavage site and determines the point at which cleavage will take place. A sequence rich in Us (or Gs and Us) is typically downstream of the cleavage site.
At the cleavage site, as many as 250 nucleotides are added following a complex process with the help some factors. (Will not be discussed here)
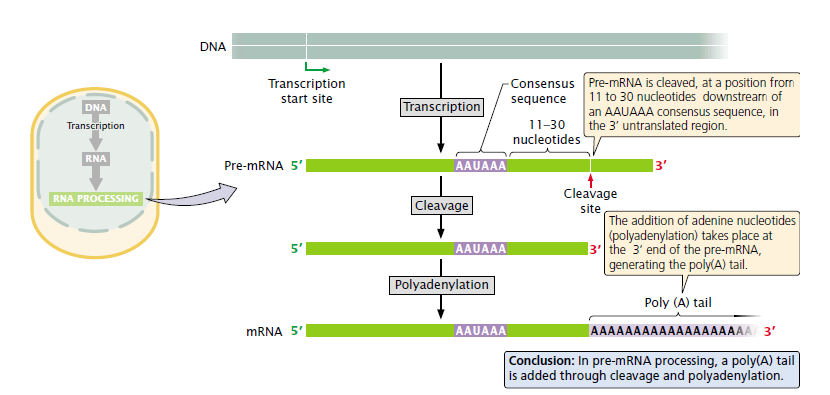
Reason: The poly(A) tail confers stability on many mRNAs, increasing the time during which the mRNA remains intact and available for translation before it is degraded by cellular enzymes.
RNA Splicing
A second important feature peculiar to the primary transcript in eukaryotes is the presence of segments of RNA, called introns or intervening sequences, that are excised from the primary transcript. Accompanying the excision of introns is a rejoining of the coding segments (exons) to form the mRNA molecule. The excision of the introns and the joining of the exons is called RNA splicing.
Splicing occurs in the nucleus following transcription but before the RNA moves to the cytoplasm.
RNA splicing takes place in nuclear particles known as spliceosomes. These abundant particles are composed of protein and several types of specialized small nuclear RNA (snRNA) molecules ranging from 100 to 200 bases in length.
(This complex process will be followed by another article and after completion, it will be hyperlinked here)
Less importance of intron
Most introns appear to have no function in themselves. An artificial gene that lacks a particular intron usually functions normally. In those cases in which an intron seems to be required for function, it is usually not because the interruption of the gene is necessary, but because the intron happens to include certain nucleotide sequences that regulate the timing or tissue specificity of transcription. The implication is that many mutations in introns, including small deletions and insertions, should have essentially no effect on gene function, and this is the case.
Moreover, the nucleotide sequence of a particular intron is found to undergo changes (including small deletions and insertions) extremely rapidly in the course of evolution, and this lack of sequence conservation is another indication that most of the nucleotide sequences present within introns are not important.
Source:
- Genetics by Heartl Jones
- Genetics A Conceptual Approach by Pierce
Revised by Md. Siddiq Hasan on 29/07/20
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka




