DNA
Before discussing “The Replication Of Circular DNA”, first we are going to give a short introduction to DNA.
DNA or Deoxyribonucleic acid is the hereditary material present in all living organisms through which the characteristics of one generation are transmitted to the next one. It contains the genetic information that makes each species unique. This molecule can be found in all prokaryotic and eukaryotic cells, even in viruses. The configuration of the DNA molecule is so stable that it enables itself to act as a template for the replication of new DNA molecules and also for the production of RNA (Ribonucleic Acid).
DNA has a double helical structure. It is a long polymer of nucleotides. The two helical chains of DNA are bound to each other by Hydrogen Bonds formed between nitrogen bases. The four bases found in DNA molecules are Adenine(A), Cytosine(C), Guanine(G), and Thymine(T), where Adenine pairs with Thymine and Guanine pairs with Cytosine. The backbone of DNA is composed of alternating Phosphate groups and Pentose (5-carbon) Sugar residues.
What Is Circular DNA?
We’ve already known that all organisms contain DNA. Generally, all organisms are classified under two classes depending on the structure of the Nucleus of the cell-
- Eukaryotic Organisms: They have a well-developed nucleus and other membrane-bound cellular organelles in their cells such as mitochondria, chloroplast, lysosome, endoplasmic reticulum, etc. Eukaryotic cells contain 80S ribosomes. Examples: humans, plants, animals, fungi, etc.
- Prokaryotic Organisms: Instead of a developed nucleus, they have circular DNA. They are in lack membrane-bound cellular organelles. They contain 70S ribosomes. Example: Bacteria, Archaea, etc.
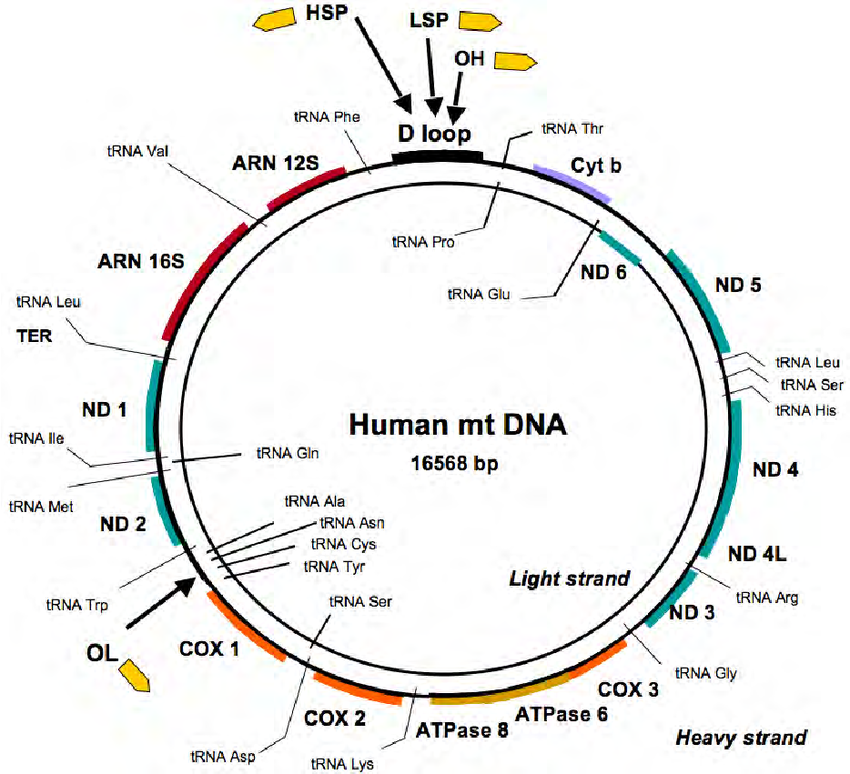
Like eukaryotes, prokaryotic organisms also have double-stranded DNA as their genetic material. But unlike eukaryotes, the DNA of a prokaryotic cell is circular. This type of DNA forms a closed loop. This type of DNA can also be found in mitochondria and plastids of Eukaryotic cells that enable these organelles to divide independently.
Circular DNA Replication
According to molecular biology, DNA replication is a biological process of producing two identical DNA molecules from one original DNA molecule.
DNA replication occurs in all living organisms which help cells to divide. It is the basis of biological inheritance.
Circular DNA replicates by Rolling Circle Mechanism. In eukaryotes, the replication of DNA is bidirectional. But the Rolling Circle Mechanism is unidirectional. The latter one is described below:
The Rolling Circle Mechanism
- Initiation
- Elongation
- Termination
Before moving into more details, first, watch the video below. (You can use the autogenerated subtitle if you face a problem understanding the audio.)
1. Initiation
- The Rolling Circle DNA replication is initiated by an initiator protein called a nicking enzyme (RepA). This protein is encoded by plasmid or bacteriophage DNA which nicks (=cuts) one strand of the DNA molecule at a site called “Double-Strand Origin” (DSO). Remember that the other strand remains as it is (no nicking).
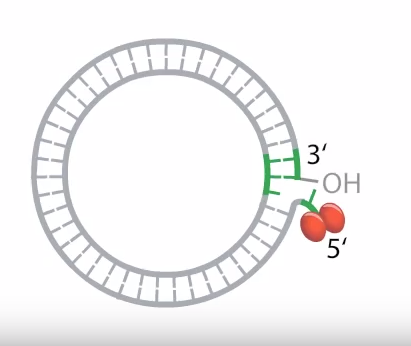
2. Elongation
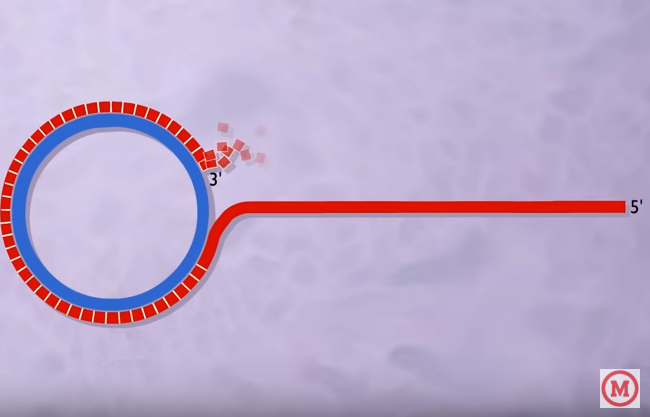
- The initiator protein remains bound to the 5`phosphate end of the nicked strand as in the figure and the 3`hydroxyl end is released to serve as a primer for DNA synthesis by the DNA polymerase enzyme.
That means the 3′ end acts as a primer.
We know that a primer is a short RNA strand used to initiate DNA replication by DNA polymerase. It is required because the DNA polymerase enzyme can’t put the complementary nucleotides in the 5′ end. It can do the replication on the 3′ end.
So, a primer is produced containing always a 5′ and 3′ end. The 3′ end acts as a point for DNA polymerase to start the replication by adding each complementary nucleotide to the 3′ end.
Now as in this case, 3′ end is already available, no additional primer is required. The nicked 3′ end acts as a primer itself.
- Meanwhile just after the nick is produced, a DNA polymerase enzyme gets attached to the complementary stand (which is not nicked or the inner circular strand) (follow figure 2).
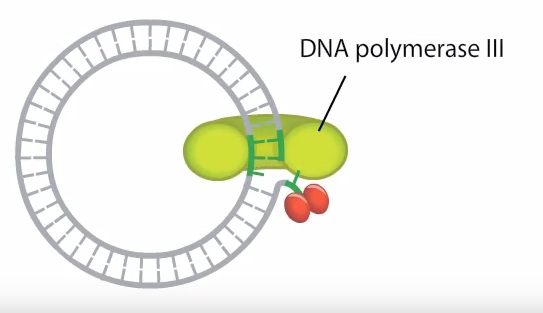
- Using the un-nicked strand (Fig 3: blue color) as a template replication proceeds, displacing the nicked strand (fig 3: the unbroken red strand) as single stranded DNA. The replication proceeds in a circular fashion. That is why it is called the Rolling Circle Model.
Good to know:
-
- The displacement of the nicked strand is carried out by a host-encoded helicase called PcrA (Plasmid Copy Reduced) in the presence of Plasmid Replication Initiation Protein.
- Continued DNA synthesis can produce multiple single-stranded DNA copies of the original DNA in a continuous Head To Tail series called Concatemer.
3. Termination
- In this step, the linear copies of the original DNA molecule are converted into circular DNA molecules.
- First, the initiator protein makes another nick to terminate the synthesis of the first (Leading) strand (the blue one). Thus the first circle is made complete.
To produce DNA from the single strand (the red one)
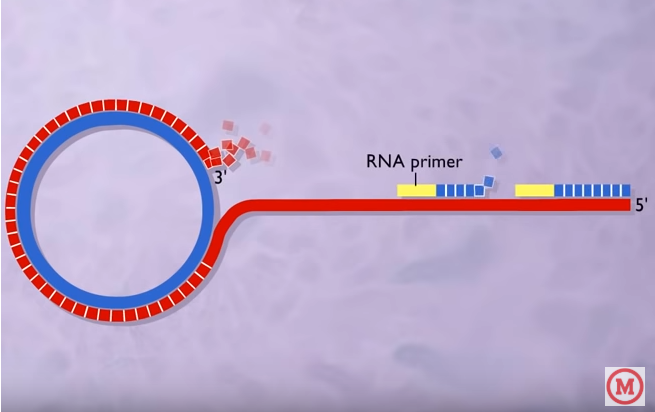
- RNA polymerase and DNA polymerase III then replicate the single-stranded origin (SSO) DNA to make another double-stranded circle.
- Then DNA polymerase I remove the primer replacing it with DNA.
- DNA ligase joins the ends making another molecule of double-stranded circular DNA.
The Rolling Circle Mechanism has been successfully used for the amplification of DNA from very small amounts of starting material.
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka




