DNA Barcoding
- DNA barcoding is a method of species identification using a short section of DNA from a specific gene or genes.
- Everyone is familiar with industrial barcodes as unique identifiers for commercial products.
- Similar to these industrial barcodes, short gene segments – known as DNA barcodes – are unique for each species.
- DNA barcoding has emerged as a global standard for fast and reliable genetic species identification of animals, plants and fungi.
A short DNA sequence of a specific gene, as for instance the mitochondrial COI gene, enables the identification of all animal species (i.e. birds, butterflies, fish, flies, etc.) on Earth. Species identification of land plants is enabled by the combination of two different chloroplast gene regions – matK and rbcL. Fungi species can be determined by the ITS region. The ultimate goal of DNA barcoding is to build a publicly accessible reference database with species-specific DNA barcode sequences. DNA barcoding is particularly effective to identify species-rich groups, cryptic species complexes, specific developmental stages (i.e. spores, eggs or larvae), and even parts and fragments of organisms (i.e. fly leg, plant root, fungal hyphae or hair). An established DNA barcode reference library offers even non-specialists a rapid, reliable and cost-effective tool for .species identification with various applications.
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
Best Money Earning Website 100$ Day
#1 Top ranking article submission website
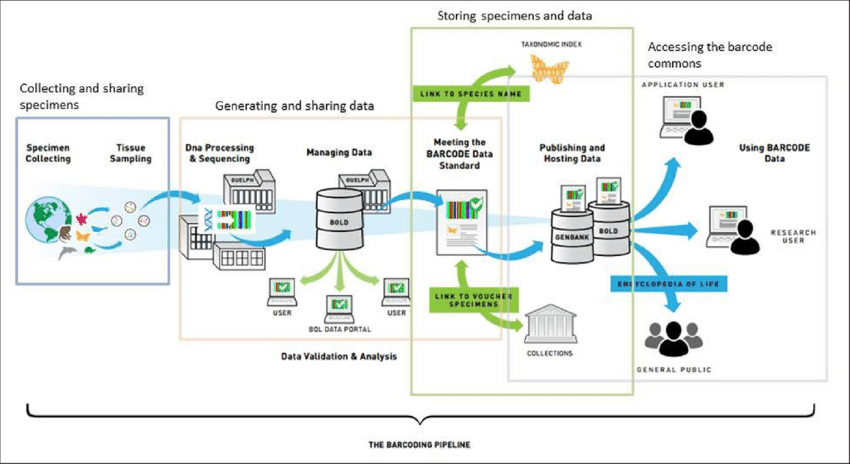
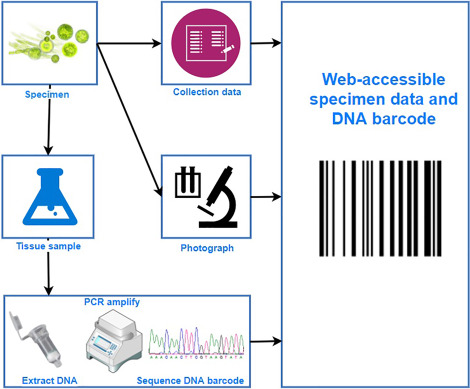
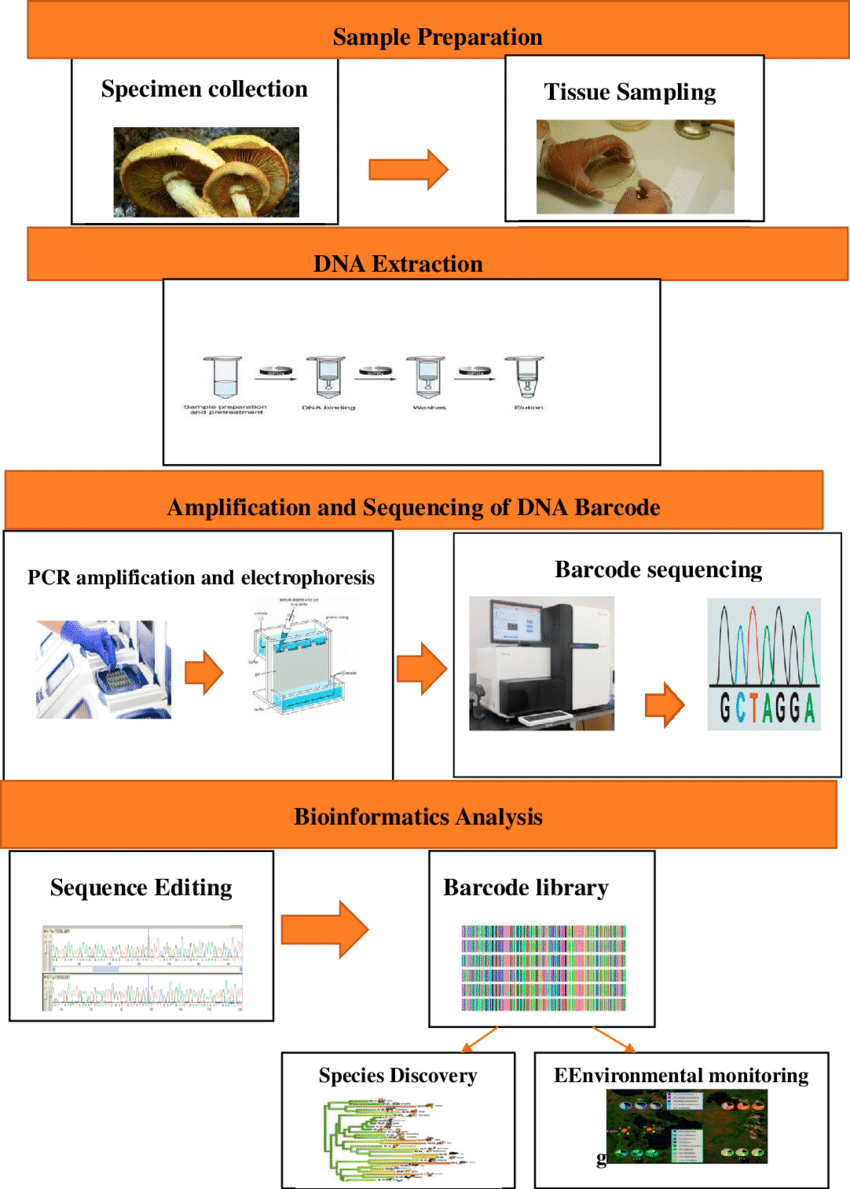
Procedure of DNA Barcoding
The process of DNA barcoding involves two basic steps: First step is to build a barcode library of identified species and second is matching the barcode sequence of the unknown sample with the barcode library (known as sequence alignment) for its identification.
- The first step requires ecologic expertise in selecting one or several individuals per species as reference samples in the barcode library.
- Tissue samples can be collected from live species in field or from specimen in museum for generating.
- These specimens go through lab processes that are tissue sampling and DNA processing and sequencing to generate DNA barcode in form of chromatogram.
- Chromatogram is visual representation of DNA sequence produced by sequencer.
- This barcode can be stored in database for future use or can be used as query sequence to be compared with sequence already present in database.
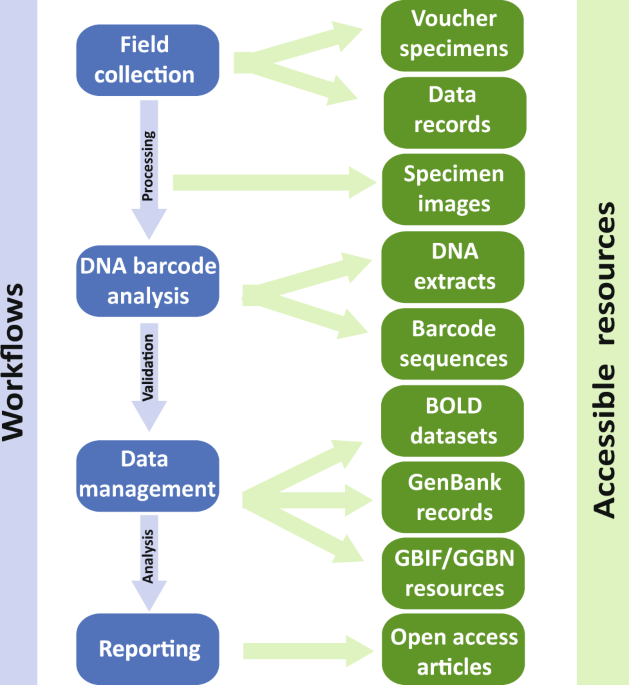
- DNA barcoding projects obtain their specimens from a variety of sources: Private collections of amateur taxonomists and museum collections (e.g. natural history museums, zoos, botanical gardens and seed banks). Museum specimens will be carefully checked for their DNA quality to ensure reliable DNA-barcoding. Fresh material collected in the field by citizen scientists. Professional GBOL taxonomists depend on the enthusiastic and active support of qualified citizen scientists. They contribute significantly to the collection, appropriate conservation and identification of organisms for DNA barcoding.
- For each sample, taxonomic identification and collection data (i.e. sampling location and date, geographic coordinates, collection method, collectors name, etc.) will be recorded.
- Organisms will be photographed in their natural habitat and in the laboratory,
- Vouchers will be stored in collections of the respectiveGBOL institutions.
- In the laboratory technicians isolate a small piece of tissue from each specimen and extract DNA.
- The DNA barcode region will be isolated and replicated using a process called PCR (polymerase chain reaction) amplification.
- The DNA barcode will be generated, consisting of a few hundred base pair long sequence of four different letters (e.g. CAATCGGTAA.). These letters represent the four nucleic acids (Adenin, Guanin, Thymin, Cytosine) constituting the DNA.
- Sequences and metadata (collection data, taxonomy, photographs) of the samples will be tested for completeness and quality by professional GBOL scientists.
- GBOL data will be uploaded to the “Barcode of Life Data Systems” (BOLD) database – an international reference library of DNA barcodes.BOLD is an online workbench that facilitates collection, management, analysis, and use of DNA barcodes.
Applications
- Applications of DNA barcoding include identification of new species, safety assessment of food, identification and assessment of cryptic species, detection of alien species, identification of endangered and threatened species, linking egg and larval stages to adult species, securing intellectual property rights for bioresources, framing global management plans for conservation strategies and elucidate feeding niches. DNA barcode markers can be applied to address basic questions in systematics, ecology, evolutionary biology and conservation, including community assembly, species interaction networks, taxonomic discovery, and assessing priority areas for environmental protection.
- Controlling Agricultural Pest: DNA barcoding can help in identifying pests in any stage of life making easier to control them saving farmers from cost of billion dollars from pest damage. The global tephritid barcoding initiative contributes to management of fruit flies by providing tools to identify and stop fruit flies at border.
- Identifying Disease Vectors: DNA barcoding allows non ecologists to identify the vector species that can cause serious infectious diseases to animals and humans, to understand these diseases and cure them. A global mosquito barcoding initiative in building a reference barcode library that can help public health officials to control these diseases causing vector species more effectively and with very less use of insecticides.
- Sustaining Natural Resources Using: DNA barcoding, natural resource managers can monitor illegal trade of products made of natural resources like hardwood trees. Fishbol is reference barcode library for hardwood trees to improve management and conservation of natural resources.
- Protecting Endangered Species: Primate Population is reduced in Africa by 90% because of bush meat hunting. DNA barcoding can be used by law enforcement to bush meat in local markets which is obtained from bush meat.
- Monitoring Water Quality: Drinking water is a process resource for living being. By studying organism living in lakes, rivers and streams, their health can be measured or determined. DNA barcoding is used to create a library of these species that can be difficult to identify. Barcoding can be used by environmental agencies to improve determination of quality and to create better policies which can ensure safe supply of drinking water.
- Routine Authentication of Natural Health Products: Authenticity of natural health products is an important legal, economic, health and conservation issue. Natural health products are often considered as safe because of their natural origin.
- Identifying of plant leaves even if flowers or fruit are not availabl.
- Identification of medical plants.
- Identification of species
- Diet analysis and food web application
- Barcoding for food safety
- Biomonitoring and ecological assessment
- Detection of invasive species
- Delimiting cryptic species
Reference
This Article is completely based on by the lecture of Dr. Mohammad Zashim Uddin, Professor, Department of Botany, University of Dhaka
Some info and pictures have been added by author
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka






The point of view of your article has taught me a lot, and I already know how to improve the paper, thank you.