What is “A Marker”?
- A marker may be defined as a “Mark of identification”.
- In Biology three major types of markers are used:
- Morphological markers (also called “classical” or “visible” markers) which are phenotypic traits.
- Biochemical markers- called isozymes-are multiple forms of enzymes, including allelic variants of enzymes.
- A genetic marker is a gene or DNA sequence with a known location on a chromosome.
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
Best Money Earning Website 100$ Day
#1 Top ranking article submission website
What is Genetic Marker?
- A genetic marker is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species. It can be described as a variation (which may arise due
to mutation or alteration in the genomic loci) that can be observed. - A piece of DNA that lies on a chromosome so close to a gene that the marker and the gene are inherited together. A marker is thus an identifiable heritable spot on a chromosome. A marker can be an expressed region of DNA (a gene) or a segment of DNA with no known coding function.
- A genetic marker may be a short DNA sequence, such as a sequence surrounding a single base-pair change (single nucleotide polymorphism, SNP), or a long one, like minisatellites.
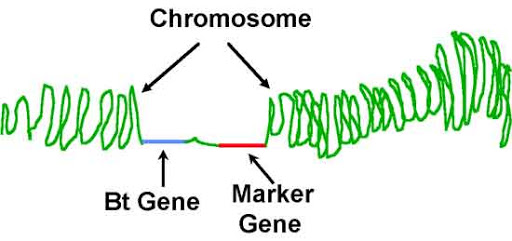
Background
- For many years, gene mapping was limited to identifying organisms by traditional phenotype markers. This included genes that encoded easily observable characteristics such as blood types or seed shapes. The insufficient number of these types of attributes in several organisms limited the mapping efforts that could be done. This prompted the development of gene markers which could identify genetic characteristics that are not readily observable in organisms (such as protein variation).
Types of genetic or molecular marker
Some commonly used types of genetic markers are:
- RFLP (or Restriction fragment length polymorphism)
- SSLP (or Simple sequence length polymorphism)
- AFLP (amplified fragment length polymorphism)
- RAPD (or Random amplification of polymorphic DNA)
- VNTR (variable number tandem repeat)
- SSR Microsatellite polymorphism, (or Simple sequence repeat)
- SNP (or Single nucleotide polymorphism)
- STR (or Short tandem repeat)
- SFP (or Single feature polymorphism)
- DArT (or Diversity Arrays Technology)
- RAD markers (or Restriction site-associated DNA markers)
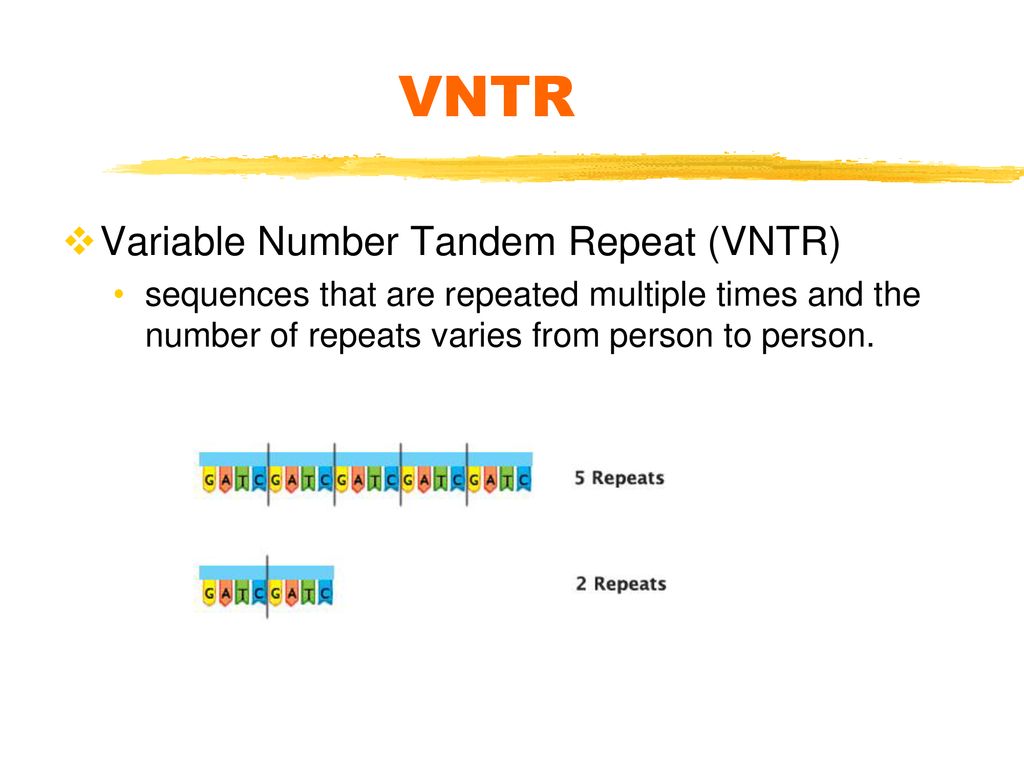
sequences that are repeated multiple times and the number of repeats varies from person to person. Another reason why every individual has a unique DNA fingerprint is because of regions called variable number tandem repeats, or VNTR’s. A VNTR is a sequence that is repeated multiple times. The number of repeats varies from person to person. This example shows the sequence GATC repeated 5 times in one individual and only 2 times in a second individual.
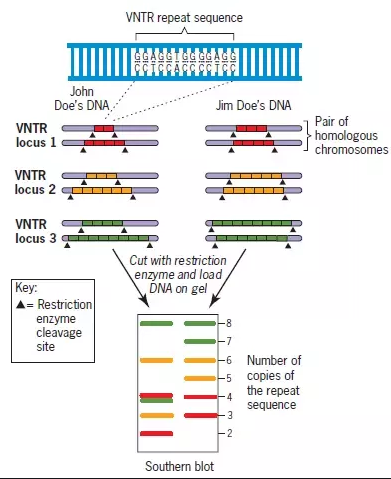
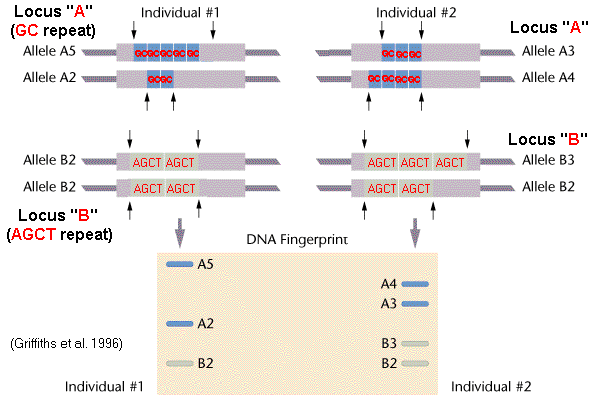
Applications genetic markers
Molecular markers have several advantages over the traditional phenotypic and biochemical markers in plant.
- The main uses of DNA markers in agricultural research such as Cultivar identity/assessment of ‘purity’/ Hybrid Testing
- to study the evolutionary relationships among individuals
- Genetic markers are employed in genealogical DNA testing for genetic genealogy to determine genetic distance between individuals or populations.
- Genetic Diversity Analysis
- Genetic Linkage Map Construction
- Mapping of quantitative trait loci (QTLs)
- Map-based cloning of genes
- Mapping of mutations
- Marker-assisted selection (MAS)
- Marker-assisted backcross breeding (MAB)
- Marker-assisted pyramiding
- Mapping major genes
- Genetic markers can be used to study the relationship between an inherited disease and its genetic cause
- Characterization of transformants
- Genetic markers can be used to study the relationship between an inherited disease and its genetic cause (for example, a particular mutation of a gene that results in a defective protein). It is known that pieces of DNA that lie near each other on a chromosome tend to be inherited together. This property enables the use of a marker, which can then be used to determine the precise inheritance pattern of the gene that has not yet been exactly localized.
- Genetic markers are employed in genealogical DNA testing for genetic genealogy to determine the genetic distance between individuals or populations. Uniparental markers (on mitochondrial or Y chromosomal DNA) are studied to assess maternal or paternal lineages, while autosomal markers are used for all ancestry.
Genetic markers have to be easily identifiable, associated with a specific locus, and highly polymorphic because homozygotes do not provide any information. Detection of the marker can be direct by RNA sequencing or indirect using allozymes. - Some of the methods used to study the genome or phylogenetics are RFLP, AFLP, RAPD, and SSR. They can be used to create genetic maps of whatever organism is being studied.
There was a debate over what the transmissible agent of CCTVT (canine transmissible venereal tumour) was. Many researchers hypothesized that virus-like particles were responsible for transforming the cell, while others thought that the cell itself was able to infect other canines as an allograft. With the aid of genetic markers, researchers were able to provide conclusive evidence that the cancerous tumour cell evolved into a transmissible parasite. Furthermore, molecular genetic markers were used to resolve the issue of natural transmission, the breed of origin (phylogenetics), and the age of the canine tumour. - Genetic markers have also been used to measure the genomic response to selection in livestock. Natural and artificial selection lead to a change in the genetic makeup of the cell. The presence of different alleles due to distorted segregation at the genetic markers is indicative of the difference between selected and non-selected livestock.
Using Antibiotic Resistance Genes As Markers
There is some concern about using antibiotic-resistance genes as markers. The antibiotic-resistance genes spread to other bacteria, producing strains of pathogenic (disease-causing) bacteria that it could not kill with antibiotics. In insulin production, the risk is probably very small, because the genetically modified bacteria are only grown in fermenters and not released into the wild. But now there are many different kinds of genetically modified bacteria around, some of which are used in situations in which their genes might be passed on to other bacteria. If these bacteria were pathogens, then we might end up with untreatable diseases.
Because of the risk of creating pathogenic antibiotic-resistant bacteria, there is now much less use of antibiotic-resistance genes in this way, and other ways have been developed in which the successfully transformed bacteria can be identified. One method uses enzymes that produce fluorescent substances. For example, enzymes obtained from jellyfish make a protein called GFP (green fluorescent protein) that fluoresces bright green in ultraviolet light. The gene for the enzyme is inserted into the plasmids. So all that needs to be done to identify the bacteria that have taken up the plasmid is to shine ultraviolet light onto them. The ones that glow green are the genetically modified ones. The same marker gene can be used in a range of organisms. Figure (A).
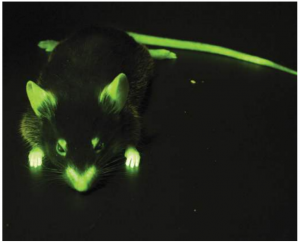
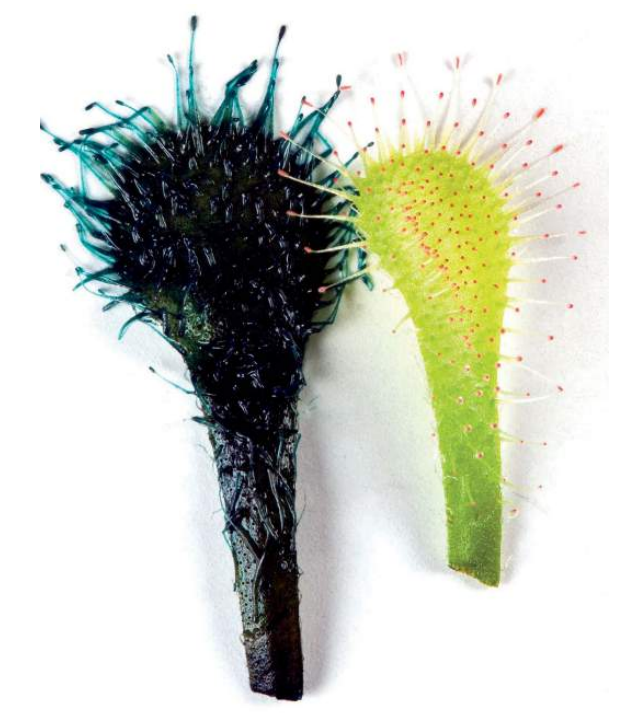
Another marker is the enzyme β-glucuronidase (known as GUS for short), which originates from E. coli. Any transformed cell that contains this enzyme, when incubated with some specific colourless or non-fluorescent substrates, can transform into coloured or fluorescent products. This is especially useful in detecting the activity of inserted genes in plants, such as the sundew
in fig (B).

So, genetic markers are useful in the identification of various genetic variations. The development of DNA-based genetic markers has had a revolutionary impact on genetic studies. With DNA markers, it is theoretically possible to observe and exploit genetic variation in the entire genome. These markers can be used to study the evolutionary relationships among individuals. Popular genetic markers include allozymes, mitochondrial DNA, RFLP, RAPD, AFLP, microsatellite, SNP, and EST markers. The application of DNA markers has allowed rapid progress in investigations of genetic variability and inbreeding, parentage assignments, species and strain identification, and the construction of high resolution genetic linkage maps for aquaculture species. The advent of next generation sequencing (NGS) has revolutionized genomic and transcriptomic approaches to biology. The new sequencing tools are also valuable for the discovery, validation and assessment of genetic markers in populations. This review focuses on the importance and uses of genetic markers with the advent of modern technologies.
Reference
This Article is completely based on by the lecture of Dr. Mohammad Zashim Uddin, Professor, Department of Botany, University of Dhaka
Some info and pictures have been added by the author
Reference used for info: Cambridge International Biology Course Book by Mary Jones
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka





