Link of other two related artciles 1 and 2. This is number 3.
(Polyploidy: A Game with Chromosome Numbers)
(Autopolyploidy: Multiplying Same Genome)
Allopolyploidy
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
Best Money Earning Website 100$ Day
#1 Top ranking article submission website
- Allopolyploids have two or more genomes derived from different genomically unlike, distinct species are present.
For example, 4x = AABB. Two types of genomes are here namely A and B. - Several of our crop plants are allopolyploids.
- Production of allopolyploids has attracted considerable attention.
Some success has been obtained as is evident from the emergence of Triticale (resulting from a cross between wheat (Triticum) and rye (Secale)) as a new crop species and Raphanobrassica (artificial hybridization of radish (Raphanus) and cabbage (Brassica) (both 2n = 18)) and some allopolyploids of forage grasses.
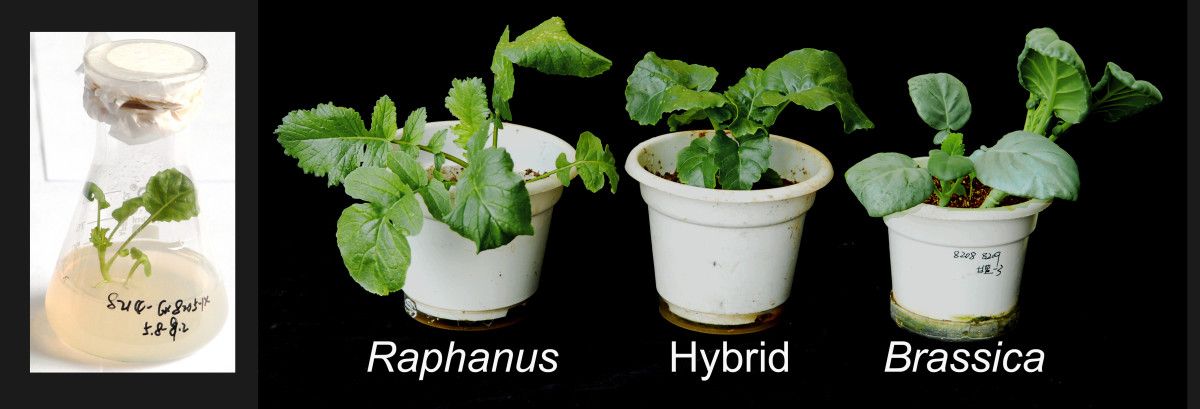

Origin and Production of Allopolyploids
- Distant Hybridization followed by chromosome doubling: The present day allopolyploids were most likely produced by chromosome doubling in sterile F1 hybrids between two genomically distinct species belonging to the same genus or two different genera.
- Irregular mitotic division in somatic cell: Chromosome doubling might have occurred in somatic tissues due to an irregular mitotic cell division leading to the formation of allopolyploid sectors either in the apical meristem or in the axillary buds. Sexual progeny from such branches would be allopolyploid.
- Production of unreduced gametes: Irregular meiosis may also lead to the production of unreduced gametes which may unite to produce allopolyploid progeny.
A complete failure of pairing may be followed by the inclusion of all the chromosomes in a single nucleus at telophase I. This would lead to the production of two unreduced gametes after telophase II. - Occassional unreduced gametes: In many distant hybrids, occasional unreduced gametes are produced which unite to give rise to allopolyploid progeny.
- Colchichine treatment in distant hybrids: Experimental production of allopolyploids is achieved by doubling the chromosome number of distant hybrids with the help of colchicine. Thus the production of an allopolyploid involves two steps: Production of F1 hybrid and chromosome doubling (producing amphiploids).
- In vitro techniques: Many distant crosses not possible previously have been successfully made with the help of in vitro techniques.
E.g. The allopolyploid Raphanobrassica was obtained from the F1 of the cross, Brassica oleracea (cabbage) x Raphanus sativus (radish) which was almost completely sterile, but produced a few seeds that gave rise to the allopolyploid Raphanobrassica.
Good to know:
- Distant F1 hybrids are generally sterile as genomes are highly divergent, and chromosomes lack affinity or form only a small number of loosely synapsed bivalents. This type of synapsis is known as heterogenetic association.
- Chromosome doubled hybrid is called amphiploid.
- The majority of allopolyploids are either tetraploids or hexaploids
Morphological and Cytological Features of Allopolyploids
Allopolyploids generally combine the morphological and physiological characteristic of the parent species. But,
- Unpredictability: It is very difficult to predict the precise combination of characters that would appear in the allopolyploid species. The interaction for any trait may be intermediiate or heterotic or lethal.
- Vigor: In general allopolyploids are more vigorous than diploids, but this also not true in all cases.
- Sterility and irregular chromosome pairing: Sterility is a common feature of amphidiploids and is most likely produced by physiological disturbances resulting from genetic imbalance. Irregular chromosome pairing, is also a contributing factor.
- Improvement of fertility: Fertility of allopolyploids can be improved through hybridization and selection.
- Highly efficient genetic system: Natural allopolyploids have evolved highly efficient genetic systems that ensure regular bivalent formation by preventing pairing between homologous chromosomes, e.g. wheat, oat (Avena sativa), tobacco etc.
- Cytoplasmic genetic male sterility: Interaction between nuclear and cytoplasmic genes may produce cytoplasmic genetic male sterility.
Good to know:
- The aim of producing Raphanobrassica was to synthesize a crop species that would combine the root of radish (Raphanus sativus 2n=2x=18; genome RR) and the leaves of cabbage (Brassica oleracea 2n=2x=18; genome BB). Raphanobrassica did combine the characteristic root and shoot systems of the two parental species, but in opposite direction.The produced F1 was an sterile one that showed absolutely no homologies. Fertility was restored by doubling the chromosomes through the formation and union of unreduced male and female gametes.
- Triticale has combineed the favourable features of the two parental species, i.e. the hardiness of rye (S. cereale) and the yielding ability of wheat (Triticum aestivum). It has great economic value. Scientists have tested the economic values of different cytotypes of Triticale particularly hexaploid (2n = 6x = 42 AABBRR) and octoploid (2n = 8x = 56)
Application of Allopolyploids in Crop Improvement
Allopolyploidy has three major applications in crop improvement :
As bridging species in the transfer of characters from one species to another
-
- E.g. Use of synthetic Nicotiana digluta (Allohexaploid) for the transfer of resistance to tobacco mosaic virus from N. sylvestris to N. tabacum. The F1 hybrid from the cross, N. tabacum x N. sylvestris is sterile. Chromosome doubling of the F1 hybrid produces the synthetic allohexaploid N. digluta which is reasonably fertile.
Production of new crop species
-
- Triticale is the most successful synthetic allopolyploids produced by crossing wheat (tetraploid or hexaploid) with rye. At present, triticale are being grown commercially in some parts of the world, e.g. in Canada the yield of triticale are comparable to those of the best wheat varieties.
Widening the genetic base of existing allopolyploid crop species
-
- The genetic base of some natural allopolyploids may be narrow and it may be useful to introduce variability by producing new allopolyploids. E.g. The genetic variability of Brassica napus has been done by crossing B. Campestris (n = 10, AA) with B. oleracea (n = 9, CC) the parental diploid species, to produce the amphidiploid B. napus (n = 19, AACC).
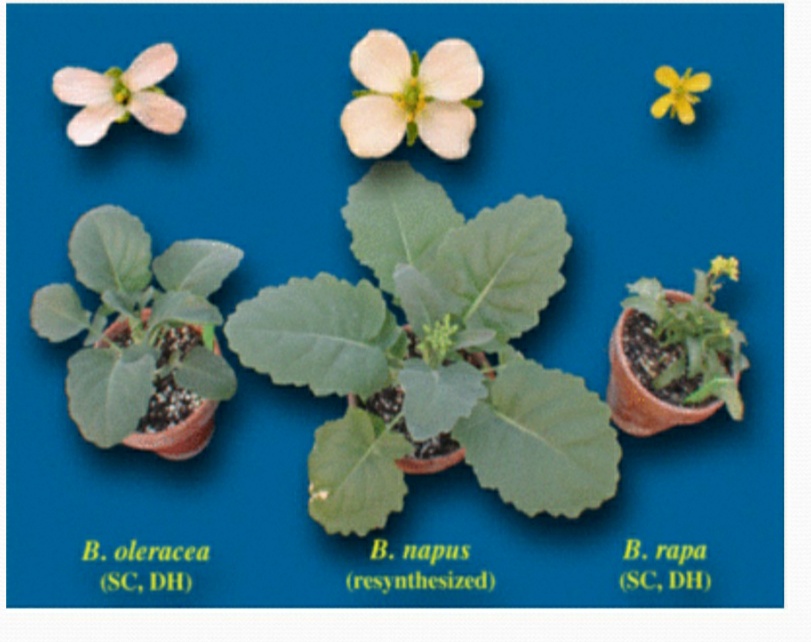
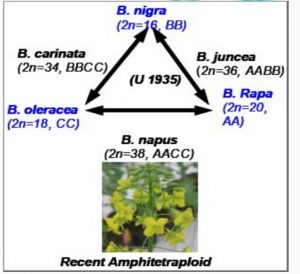

Role of Alloployploidy in Evolution
Allopolyploidy has contributed to a great extent in the evolution of plants. It is estimated that about one-third of the Angiosperms are polyploids and by far vast majority of them are allopolyploids.
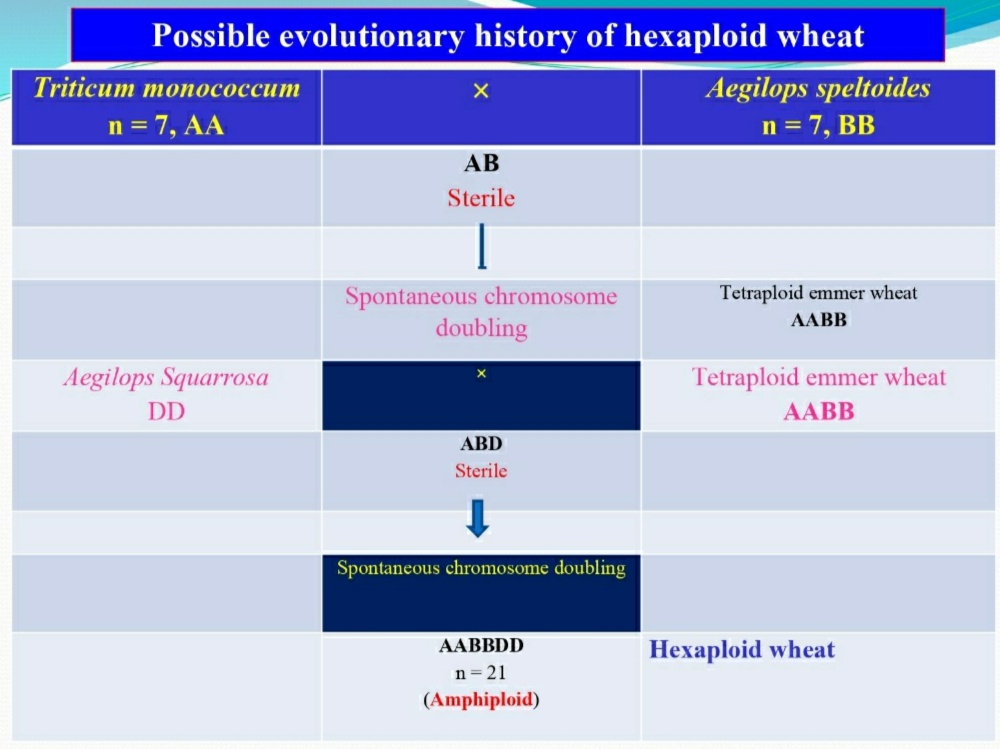
Evolution of Bread Wheat (Triticum aestivum):
- Evolutionary history of wheat has been the most extensively investigated and is perhaps the least understood.
- Identity of the diploid species contributing the three genomes (A,B and D) of T. aestivum has been investigated by many workers (Sears, Kihara and others).
- It has been proposed that the source of ‘A‘ genome is T. monococcum, ‘B‘ genome is Aegilops speltoides and ‘D‘ genome is Aegilops squarrosa.
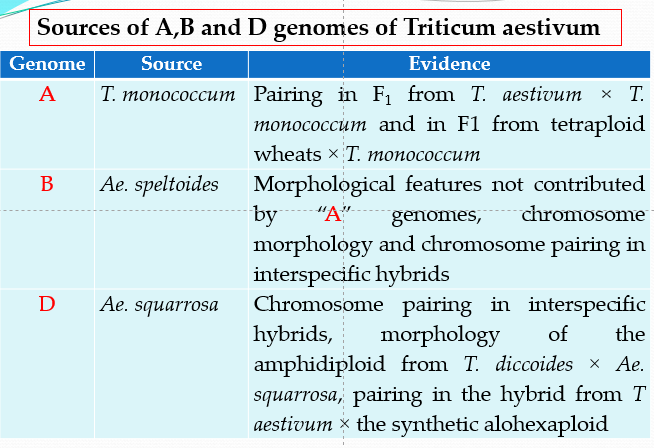
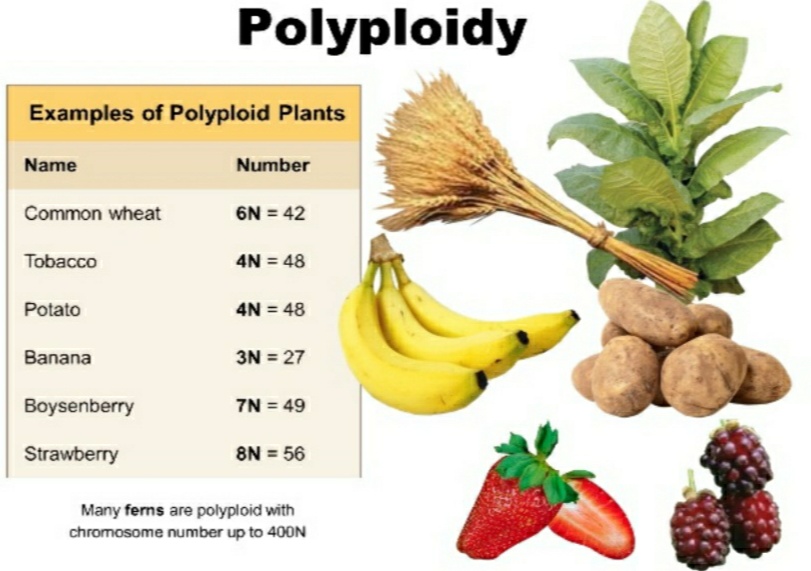


Source:
Featured image: Bioninja – https://ib.bioninja.com.au/higher-level/topic-10-genetics-and-evolu/103-gene-pools-and-speciati/allopolyploidy.html
Best safe and secure cloud storage with password protection
Get Envato Elements, Prime Video, Hotstar and Netflix For Free
 Plantlet The Blogging Platform of Department of Botany, University of Dhaka
Plantlet The Blogging Platform of Department of Botany, University of Dhaka





The point of view of your article has taught me a lot, and I already know how to improve the paper, thank you.